During my stay in Paris, one of the topics I have been working on is related to the problem of imaging objects hidden by scattering media. While seeing stuff through transparent materials is quite easy, the thing becomes much more complicated when light gets randomly deviated in all directions. This might seem quite a particular scenario, but basically any material we have around scatters light to some degree. In fact, in most of the interesting imaging applications, scattering is one of the biggest problems that optical systems need to face. No matter if you are looking at the stars with a telescope or inside a mouse with a microscope: your images will rapidly become blurred due to the rapid changes in the atmosphere or if light has to travel through more than a few hundred microns of tissue. This effect can be seen in the following picture:
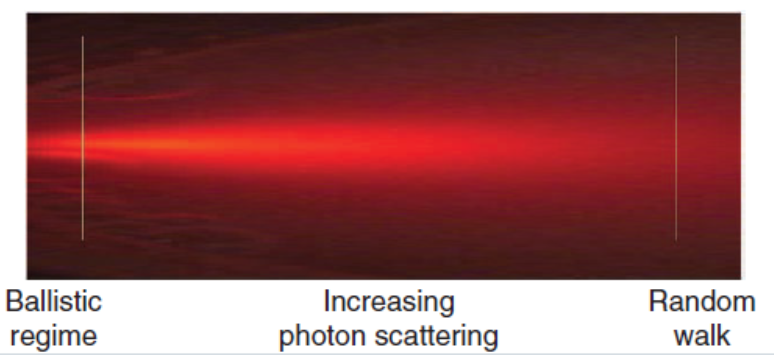

It is easy to see that at low penetration depths, light keeps its original direction. However, when going inside the scattering media, light spreads out in random directions, and after just a bit of material you get a diffuse halo of light that cannot be used to obtain an image. Or is it? Actually, if you try to focus coherent light through a highly scattering sample, you will get a totally random, seemingly information-less speckle pattern instead.
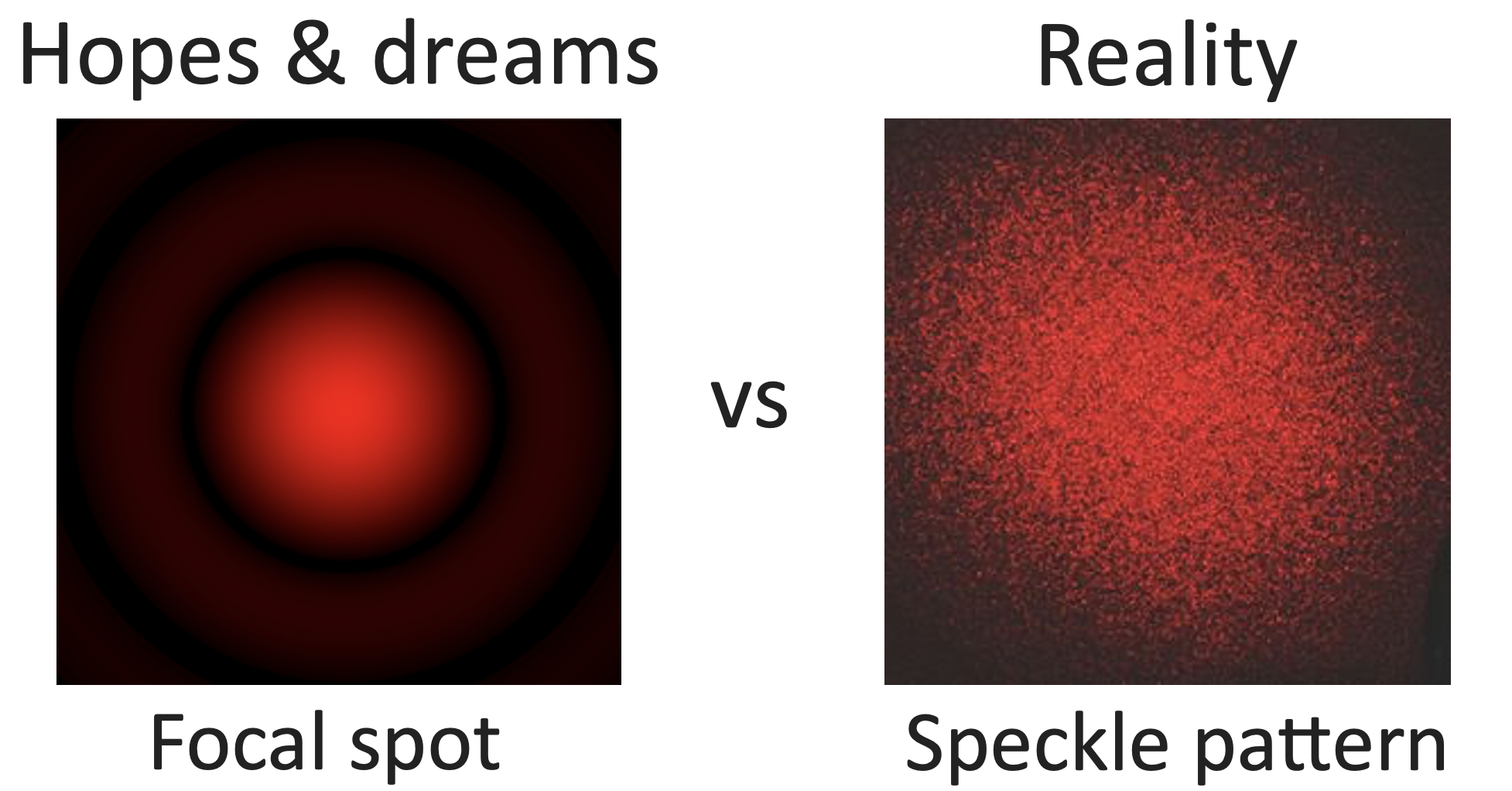

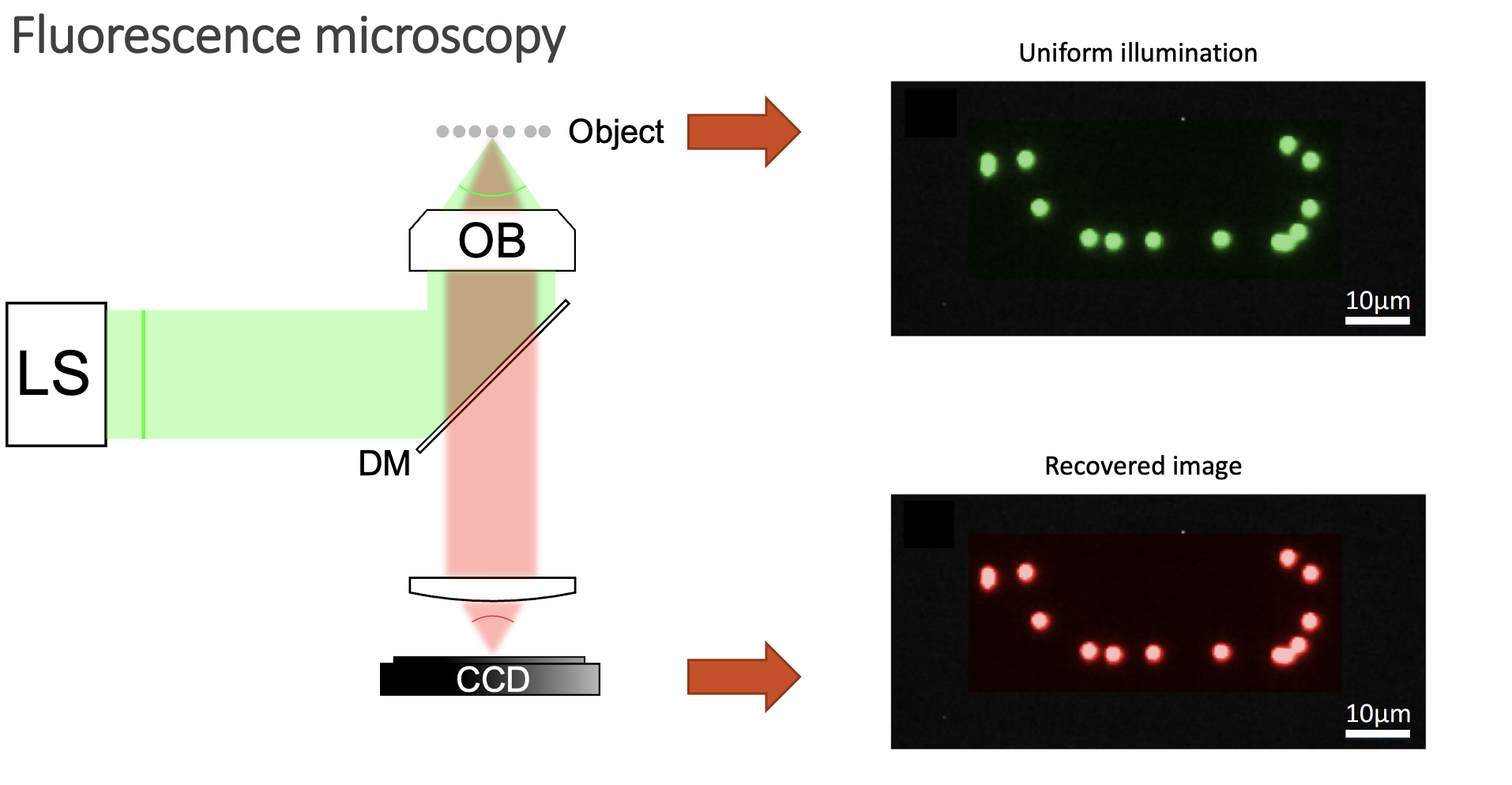
This makes common microscopy setups unable to obtain good images. Usually, you want to deliver light uniformly inside your sample, and then image your object.
However, when you have a scattering medium, things get a bit more complicated:
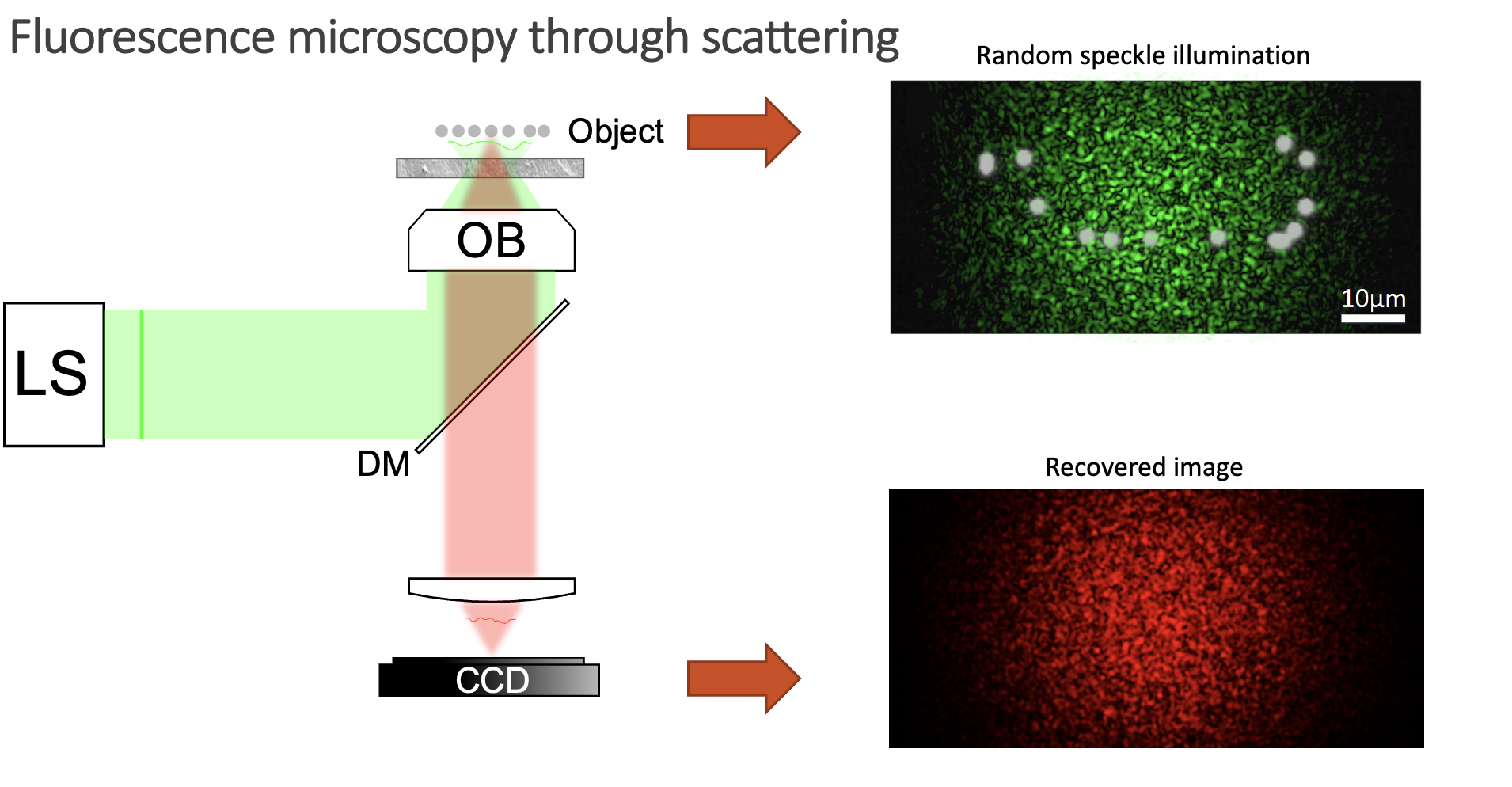
Now, instead of illuminating the whole sample, you blindly excite fluorophores with a speckle pattern. Some emitters will be excited, but others will lay on dark regions of the speckle pattern and thus will not emit any signal. Moreover, the fluorescent signal also gets scrambled on its way out of the sample, so your detector can only obtain a speckle pattern and not a sharp image. So, is there anything we can do to retrieve the image of the object in this case? Well, it might seem like a very difficult task, as we do not know how we are illuminating the sample, and the signal gets scrambled on its way out. However, there are a few things we know about the system:

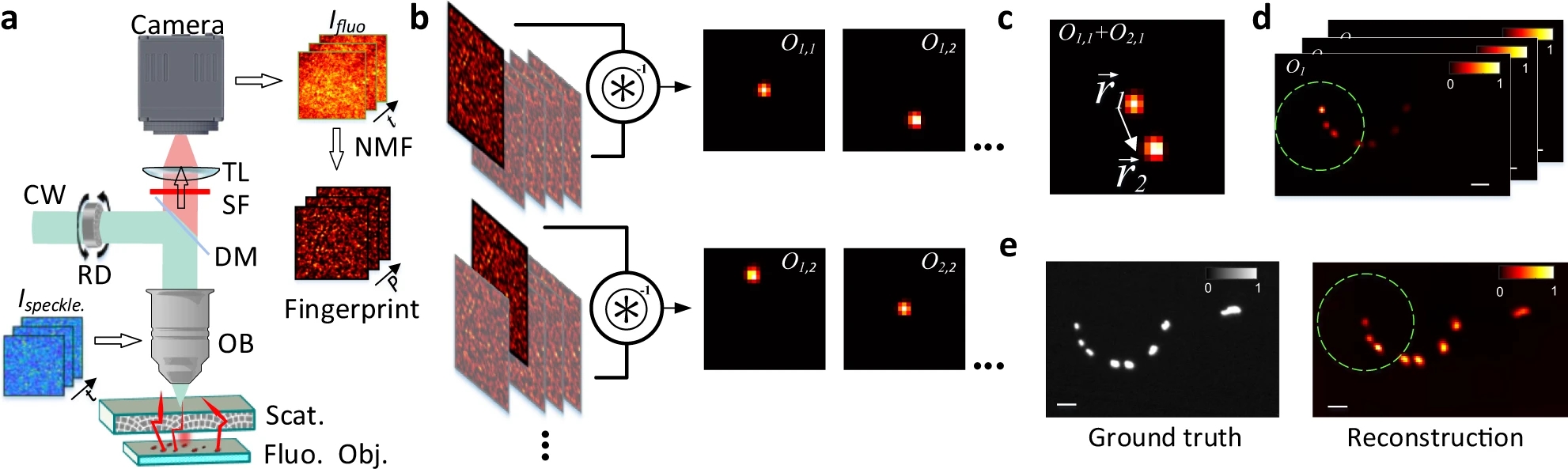
We know that, while random, there is an excitation pattern onto the object, so it is quite probable that we will excite most of the emitters. Also, each emitter will generate a unique speckle pattern onto the detector, and those will be added together. In other words, we can express the image we capture as a linear combination of a set of unknown speckle patterns. Furthermore, if the emitters are close, the memory effect establishes that their speckle patterns will be highly correlated, but laterally shifted versions of each other. If we get access to all the individual speckle patterns (we could think of them as fingerprints), we should be able to retrieve the relative position between all the emitters.
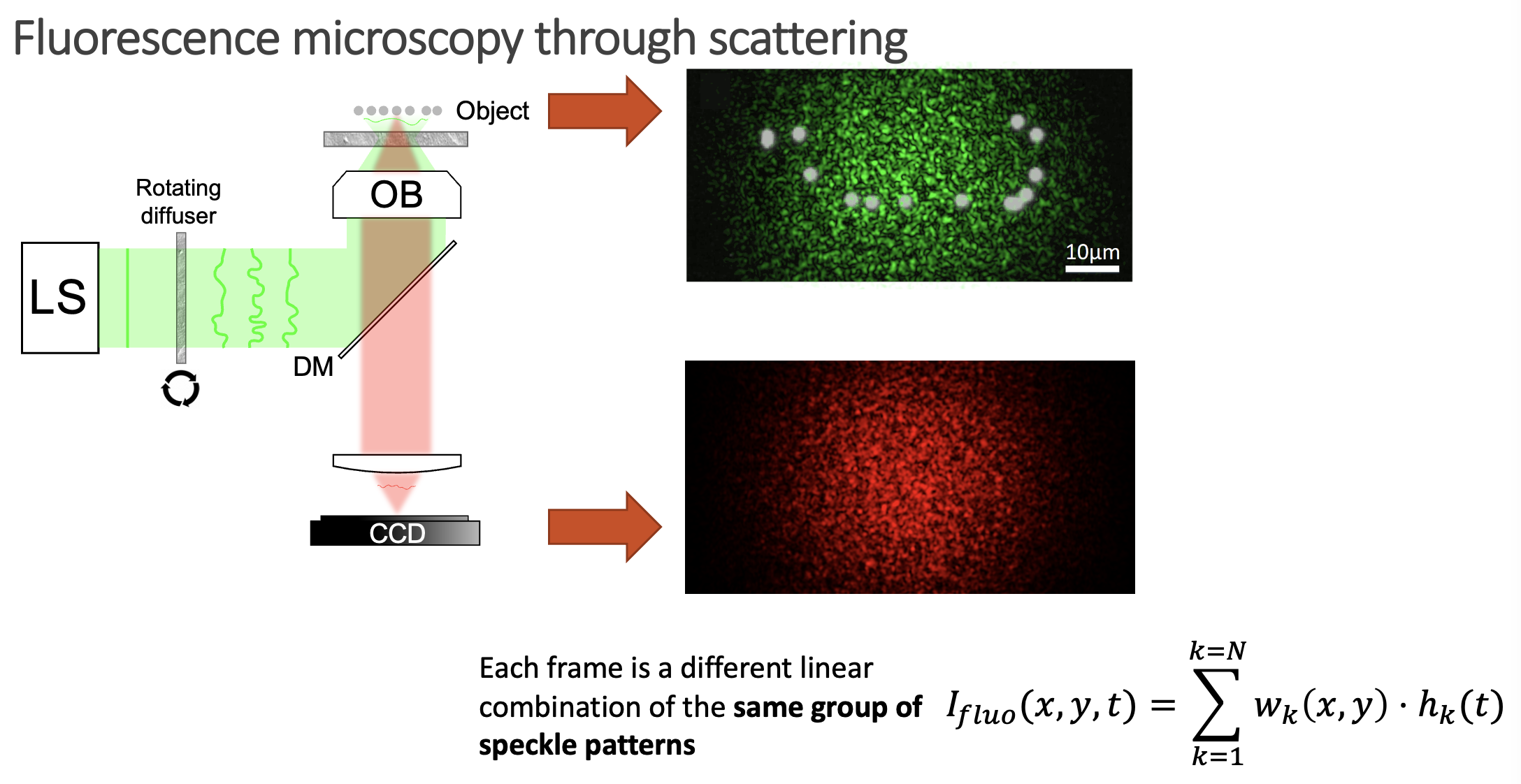
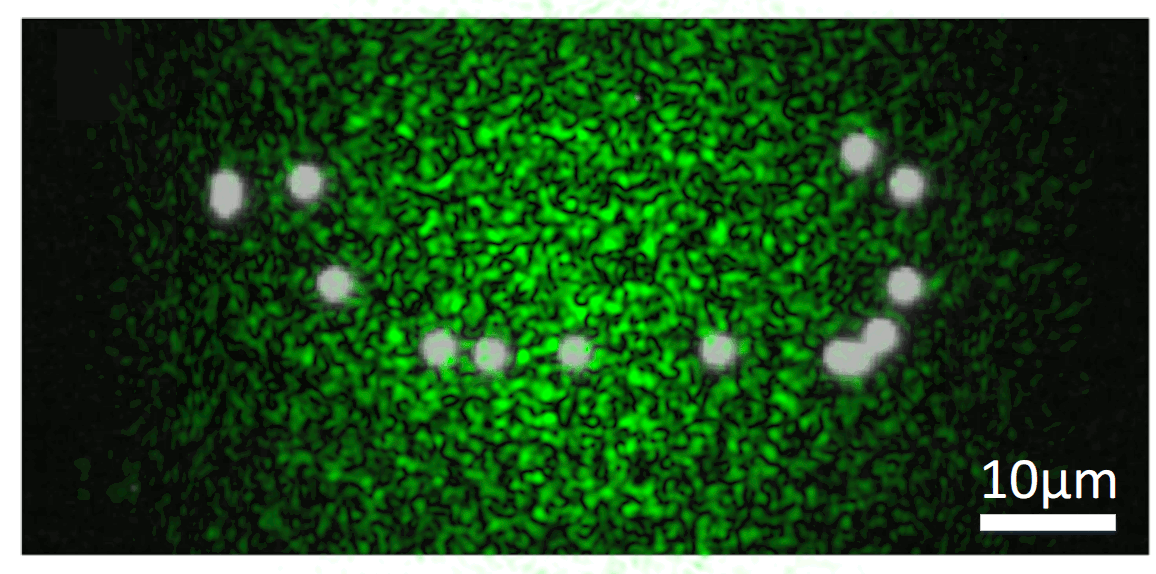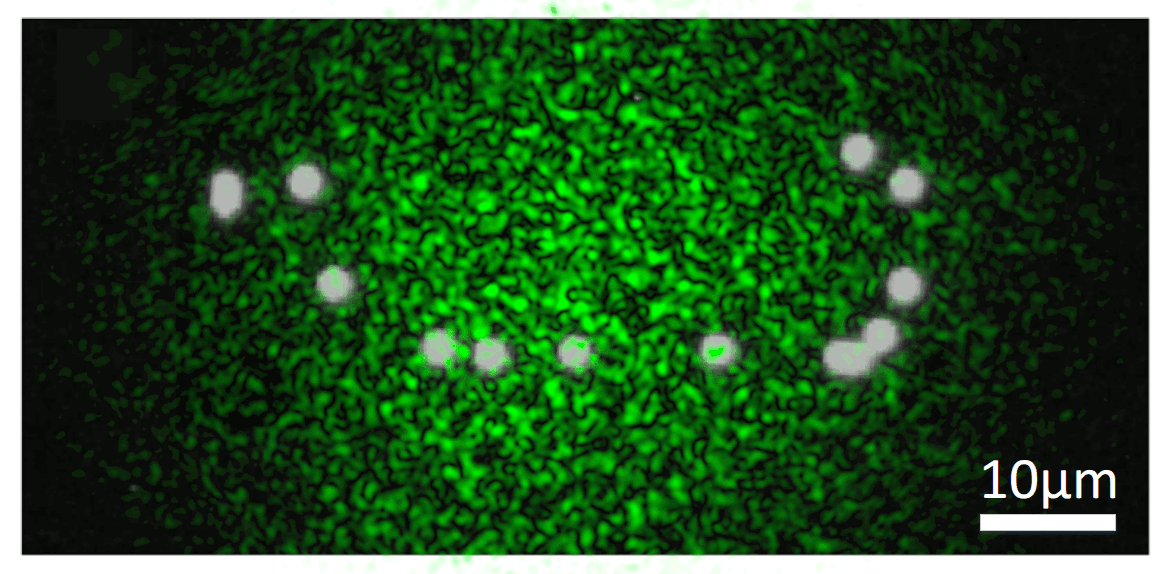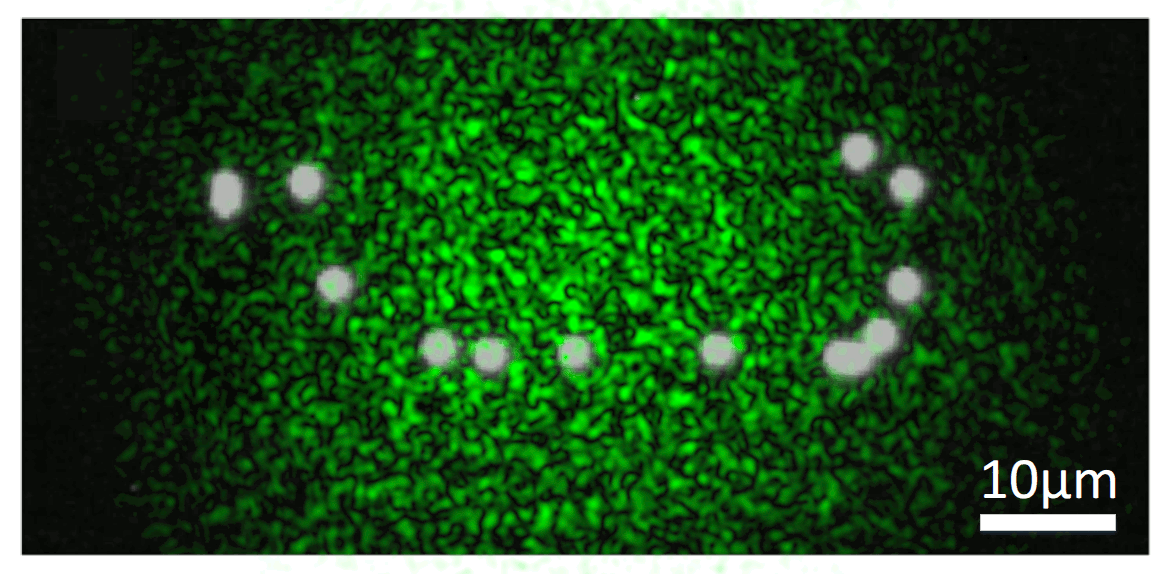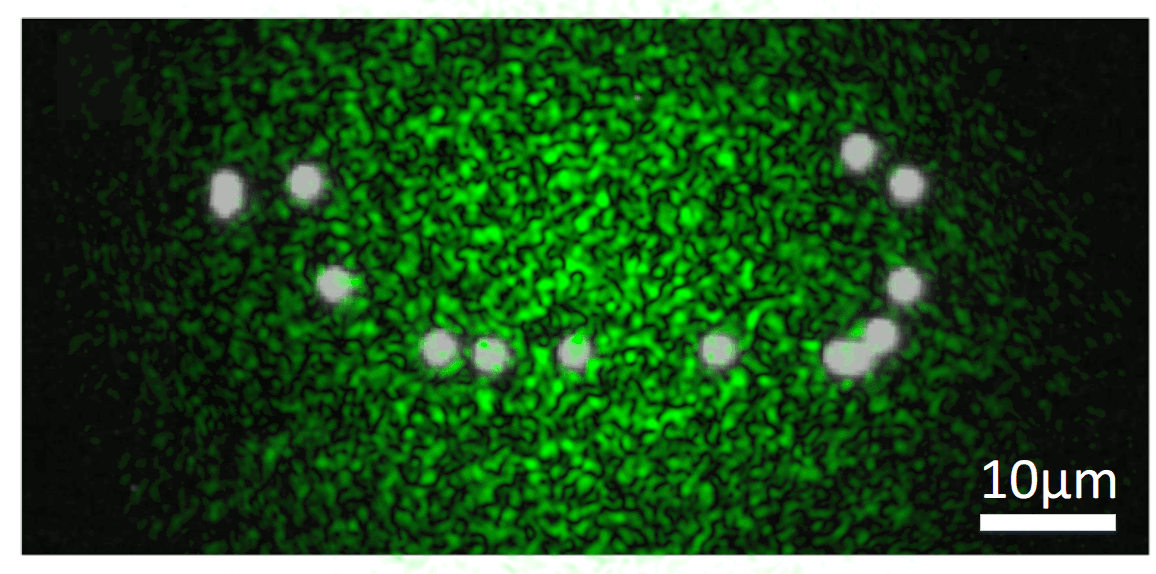
But, it is really possible to get all the individual speckles? Well, that is a tricky question. From a single shot, it is not. However, if you change the illumination over time, your detector will record different linear combinations of the same reduced library of speckle patterns. Then, if you gather enough of these different linear combinations (each linear combination will have different weights on the fingerprints, but the fingerprints will always be the same), it is possible to demix them all by solving an optimisation problem. This we can experimentally achieve in a simple manner by introducing a rotating diffuser, which will change the speckle illumination over time.

The unmixing problem is commonly known as Nonnegative Matrix Factorization (NMF), as the whole video we acquire can be expressed as a big matrix which entries are positive (or zero), and we want to retrieve the two positive matrices that represent it (W containing the speckle fingerprints, and H storing the temporal activity information).
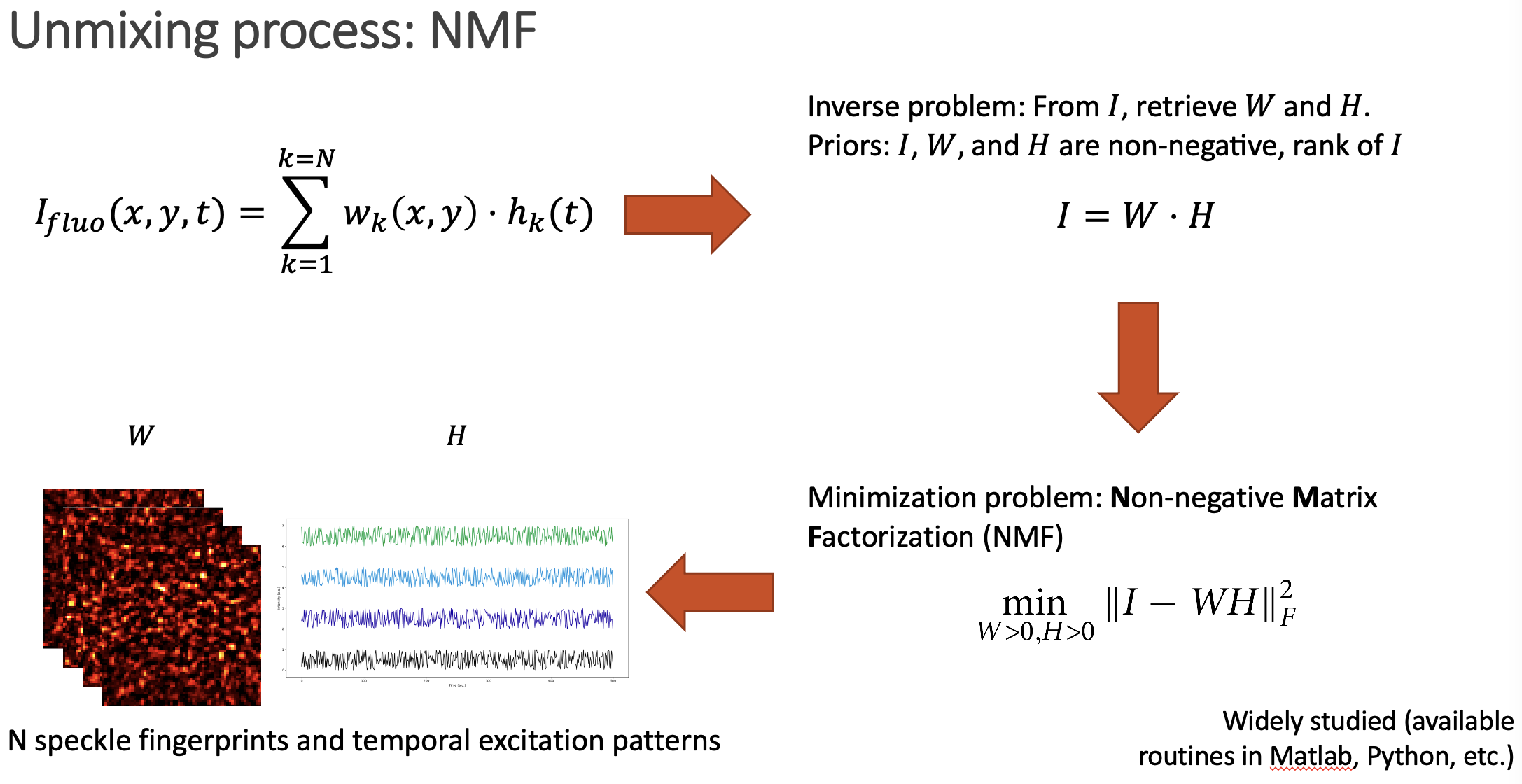
After solving the NMF problem, we get access to the individual speckle patterns that each emitter generates. Then, it is just a matter of comparing them to retrieve the spatial distribution of emitters.
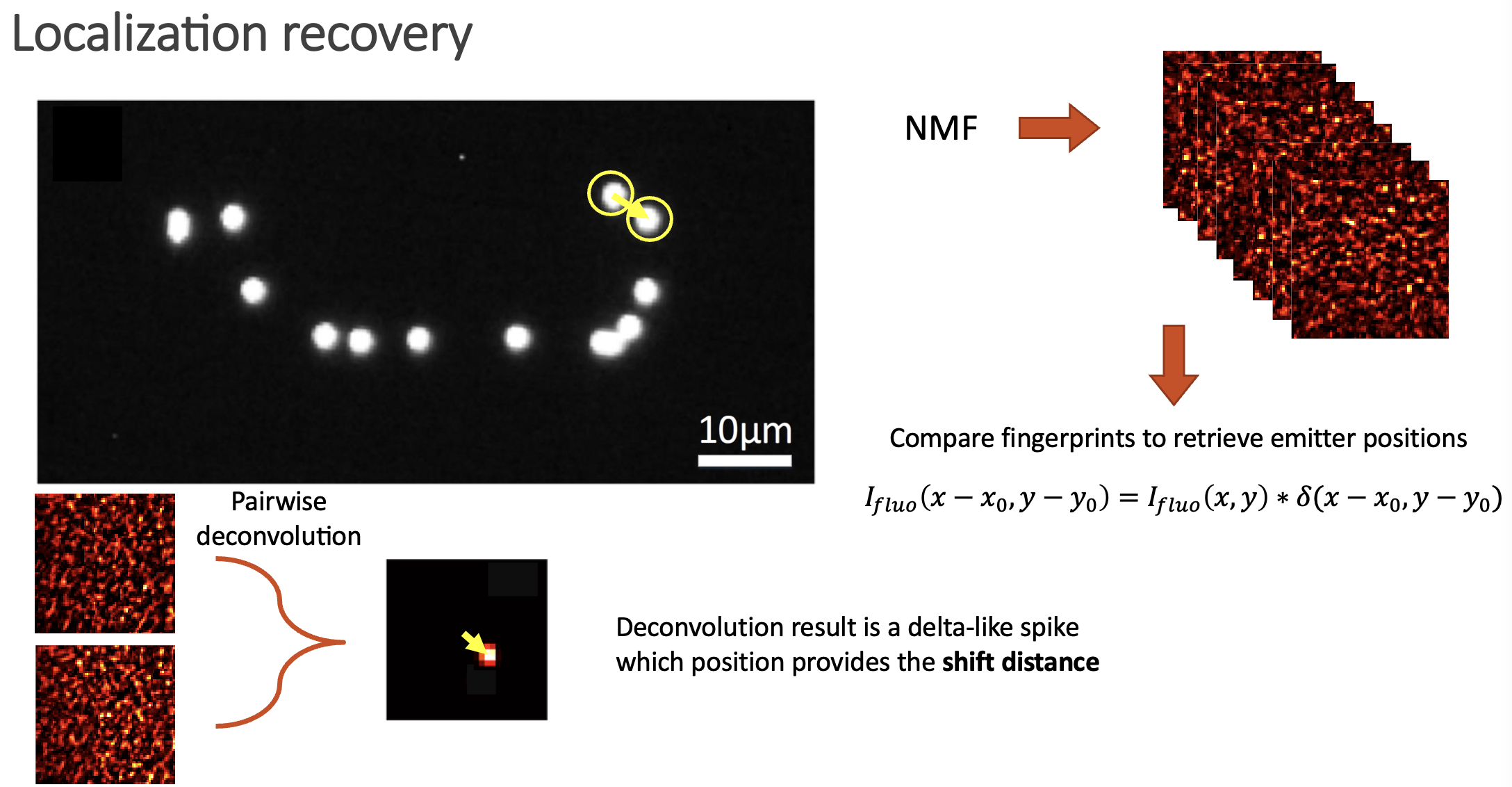
If two speckle patterns come from neighbouring emitters, they will be highly correlated (but laterally shifted). We can always link two shifted images via a convolution with a Delta function. Then, if we deconvolve two speckle fingerprints, the result should be an image with a delta spike, which location provides the spatial shift between the two images (if the images are highly correlated), or just noise if the images are not similar.
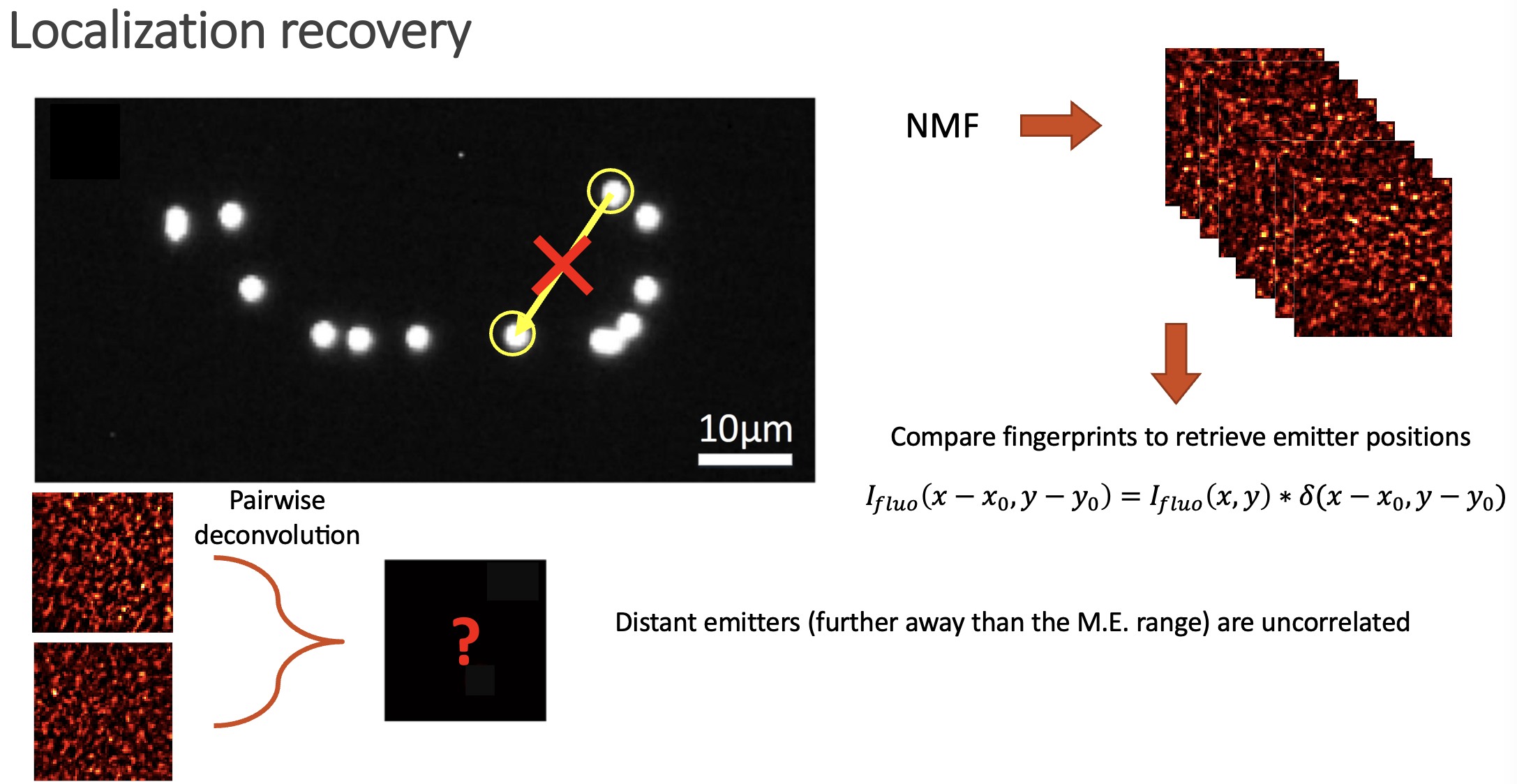
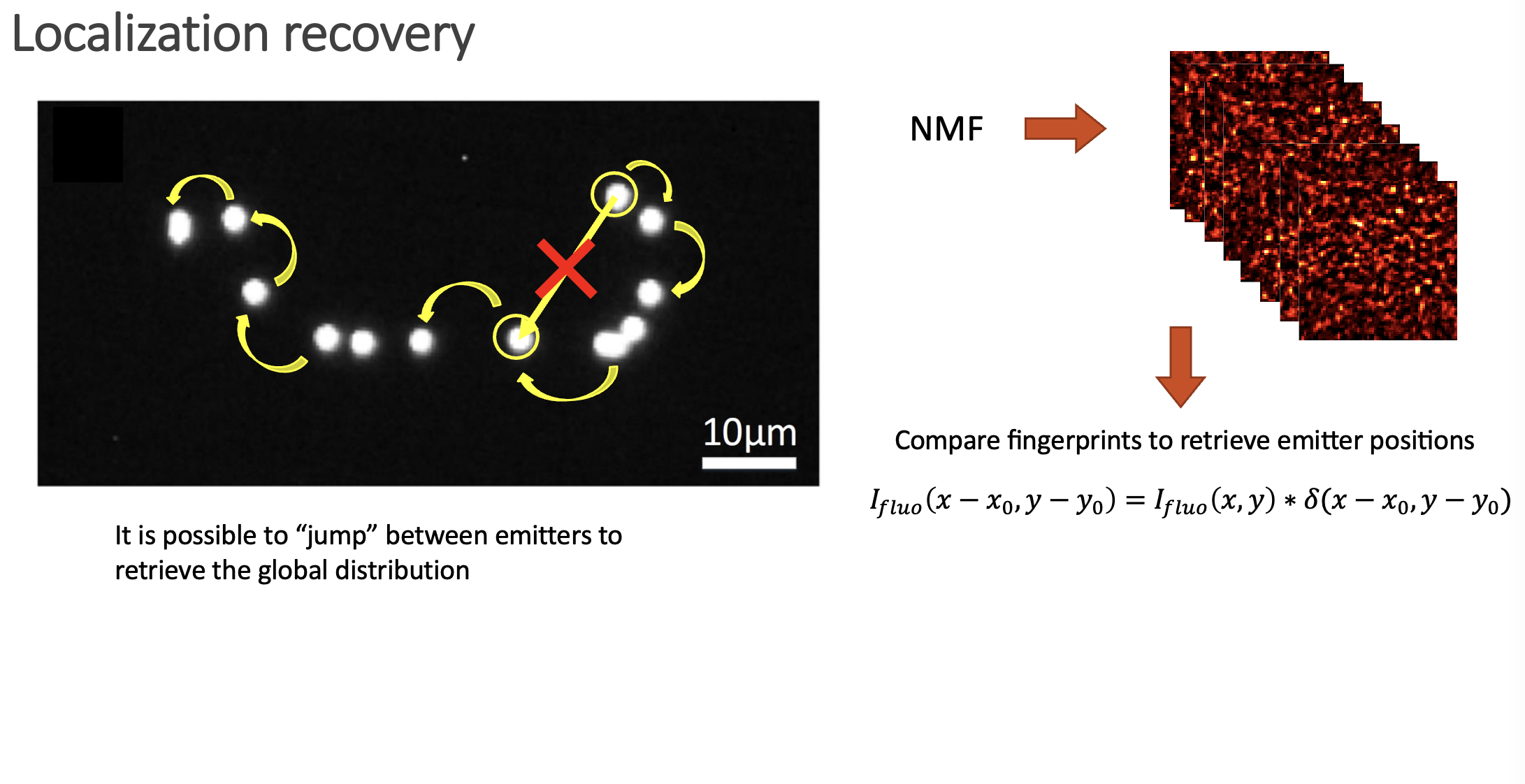
In the paper we show how is possible to deconvolve all the possible combinations between pairs of fingerprints, and then build a location map by jumping between neighbouring emitters. Even if the emitters span over large distances, the whole picture can be retrieved.
This procedure allows to retrieve both sparse objects (bead distributions), but also continuous objects (pollen grains and cellulose fibers) over large fields of view:
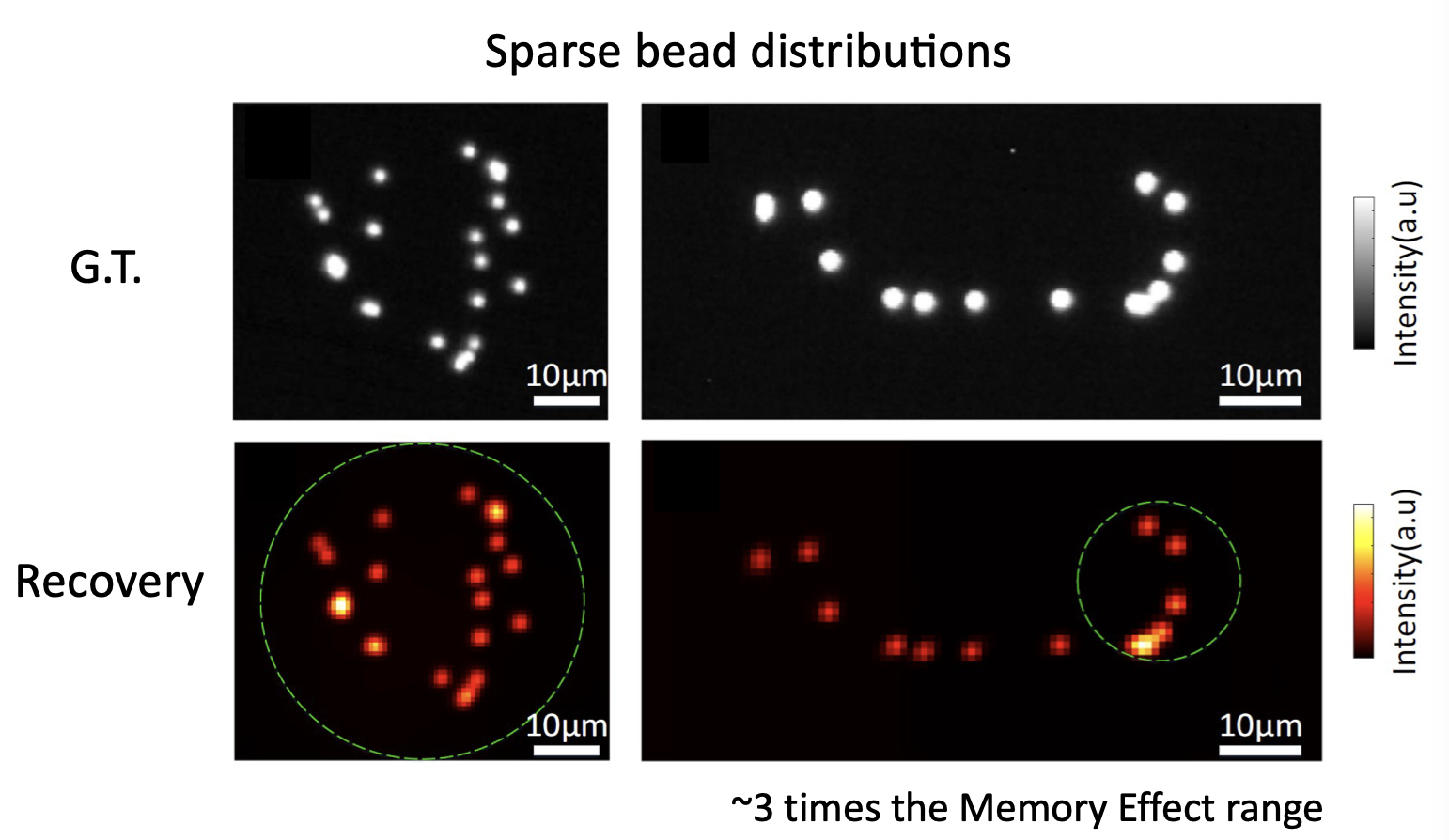

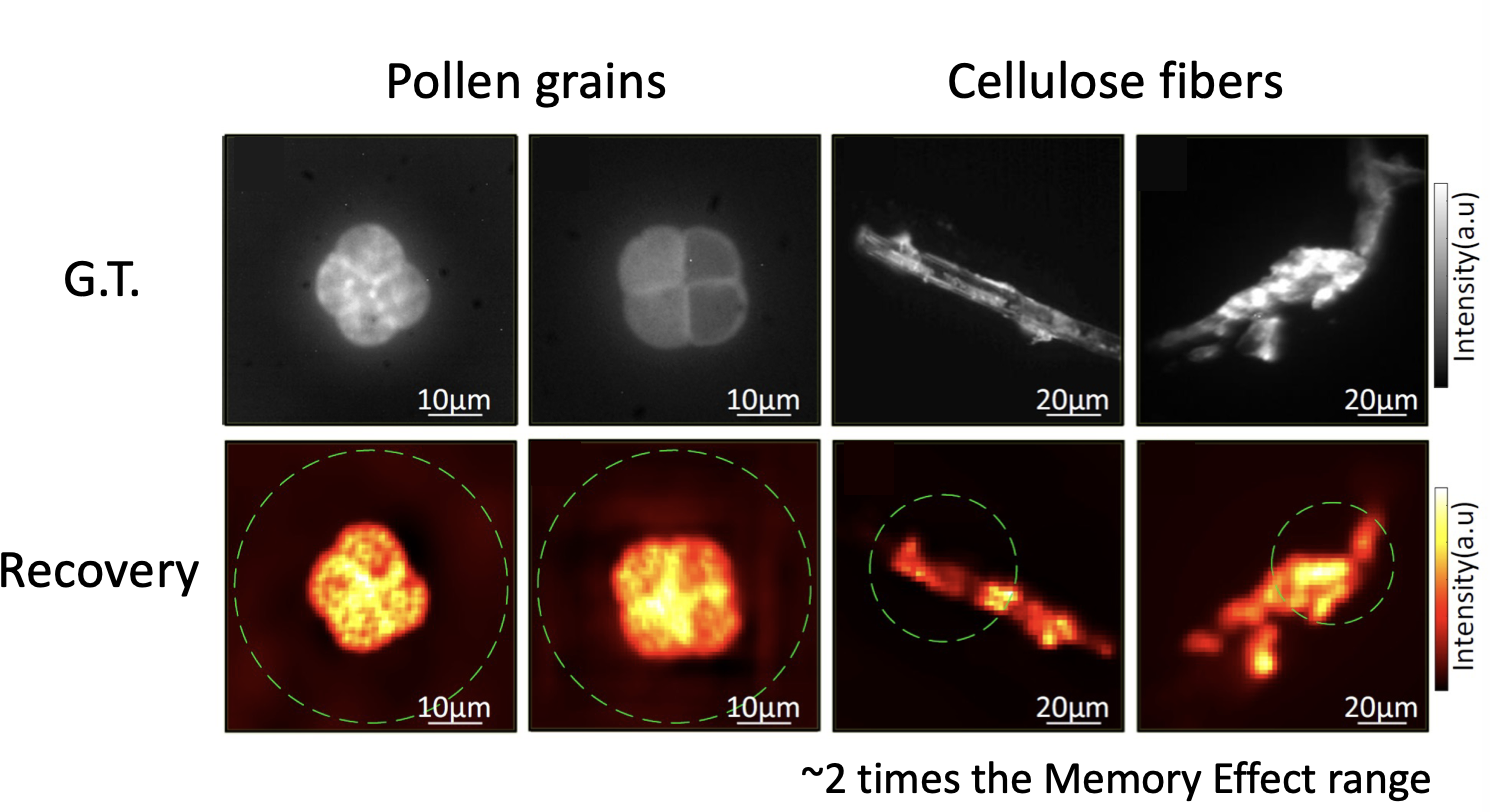
The cool thing about this approach is that is totally non invasive. You do not need to characterise your medium at all (even if you do not know how your sample is being illuminated, you can still retrieve its shape). Moreover, the optical system is quite simple, as you only need to use a rotating diffuser to be sure that the illumination is changing over time. From the computational point of view the system is quite appealing, as NMF can be efficiently solved, and there is a lot of room of adding any a priori knowledge you might have about your system (sparsity, etc.). You can find all the details about the technique in the manuscript, and the codes/blueprints to make it work on the Github repo.
Large field-of-view non-invasive imaging through scattering layers using fluctuating random illumination
Lei Zhu, Fernando Soldevila, Claudio Moretti, Alexandra d’Arco, Antoine Boniface, Xiaopeng Shao, Hilton B. de Aguiar & Sylvain Gigan. Large field-of-view non-invasive imaging through scattering layers using fluctuating random illumination. On Nat Commun 13, 1447 (2022). https://doi.org/10.1038/s41467-022-29166-y
Abstract:
Non-invasive optical imaging techniques are essential diagnostic tools in many fields. Although various recent methods have been proposed to utilize and control light in multiple scattering media, non-invasive optical imaging through and inside scattering layers across a large field of view remains elusive due to the physical limits set by the optical memory effect, especially without wavefront shaping techniques. Here, we demonstrate an approach that enables non-invasive fluorescence imaging behind scattering layers with field-of-views extending well beyond the optical memory effect. The method consists in demixing the speckle patterns emitted by a fluorescent object under variable unknown random illumination, using matrix factorization and a novel fingerprint-based reconstruction. Experimental validation shows the efficiency and robustness of the method with various fluorescent samples, covering a field of view up to three times the optical memory effect range. Our non-invasive imaging technique is simple, neither requires a spatial light modulator nor a guide star, and can be generalized to a wide range of incoherent contrast mechanisms and illumination schemes.
Deja una respuesta